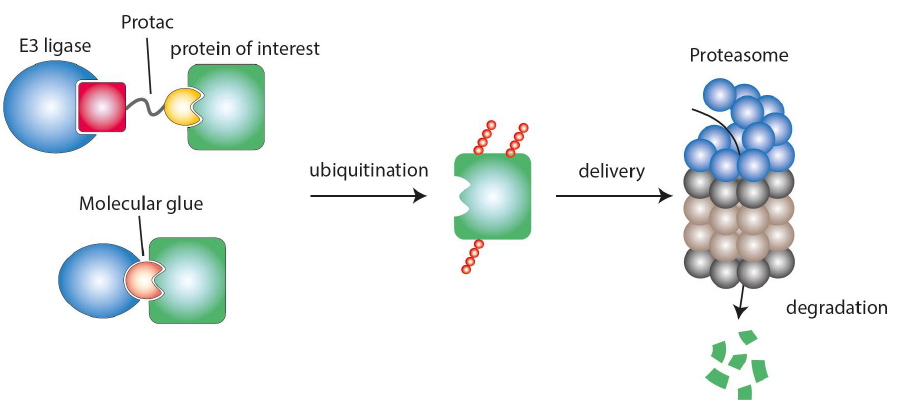
The aim of the platform is to utilize the proximity-inducing drugs (PiDs) class of active ingredients for the treatment of immune-mediated diseases. PiDs are bifunctional molecules that transiently bind to two target structures, bringing them into proximity and thereby triggering a biological effect. In the case of PiDs in the narrower sense, one of the two structures is an ubiquitin E3 ligase, the other is the target protein, which is labelled by this E3 ligase for degradation in the proteasome. These PiDs thus lead to the complete loss of the target protein with all its catalytic, structure-giving or regulatory functions, which distinguishes PiDs from classical inhibitors.
80% of the human proteome is considered ‘undruggable’
Despite a large arsenal of available drug classes, about 80% of the human proteome is considered therapeutically inaccessible or difficult to access. An active site is not required for the effect of PiDs – this makes it possible to address disease-related target proteins that were previously considered undruggable.
PiDs, in particular PROTACs (proteolysis-targeting chimeras), are a new but very intensively researched class of molecules. The most advanced agents in this class are currently in the early phases of clinical development, primarily for oncological diseases.
Characterization and optimization of PiDs
The work within the platform is intended to establish the infrastructure, processes and workflows for the design, synthesis, characterization and optimization of PiDs in a cross-institutional collaboration. Two classes of PiDs will be addressed: PROTACS and LYTACs (lysosomal-targeting chimeras). Using one or more selected target proteins as examples, PiDs will be identified, their effectiveness tested in models of immune-inflammatory diseases and their mode of action compared with that of classic inhibitors.
 Fraunhofer Cluster of Excellence Immune-Mediated Diseases
Fraunhofer Cluster of Excellence Immune-Mediated Diseases